Yeast Mutant Repository Under Construction
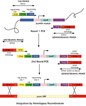
Researchers in the consortium, known as the Saccharomyces Genome Deletion Project, are systematically deleting every yeast gene using a PCR-mediated gene disruption strategy that also inserts a barcode-like tag to provide a permanent marker for each strain. Eventually, they will end up with 6,000 different yeast strains, each one lacking a single gene. Laboratories interested in studying a particular gene will no longer have to invest the individual time and effort into making the deleted yeast strain. They can simply request their strain of choice.

So far, 6,925 mutant yeast strains have been constructed which collectively contain deletions in more than one third of the yeast open reading frames. And already the analysis of these strains has led to some interesting and important findings:
- 17% of the deleted genes are essential for survival even when grown in rich medium.
- 40% behaved as conditional lethals.
- 8.5% of non-essential genes have homologs within the yeast genome, perhaps accounting for their not being essential for growth.
- Essential genes tended to be clustered and more heavily transcribed than non-essential genes.
The presence of a unique molecular tag in deletion strains provides an advantage over other methods for creating deletions: it allows individual mutants to be identified within a complex mixture so that many different yeast can be tested at once. More than 500 mutant yeast can now be analyzed in parallel, greatly increasing speed, accuracy, and efficiency.
According to Elizabeth Winzeler, lead author of the article and a postdoctoral fellow in the laboratory of Stanford biochemistry professor Ronald Davis, it is theoretically possible that a group of yeast containing deletions in every gene in the genome could be tested at the same time. "Once the set of deletion strains is completed, function might be assigned to many genes using a hundred or so different experiments, instead of hundreds of thousands of experiments," she said.
Winzeler and her colleagues collaborated with researchers at Affymetrix Inc. (Santa Clara, CA) to analyze the yeast genome and design the molecular tags and other reagents specific for each strain. The materials were then sent to consortium members who made the deletions. The final products are being distributed to the wider research community via public repositories in Europe and the United States.
The study was conducted in collaboration with researchers from Johns Hopkins University School of Medicine (Baltimore); McGill University (Montreal); Rosetta Inpharmatics Inc. (Kirkland, WA); Affymetrix; Yale University (New Haven, CT); Washington University Medical School (St. Louis); FYSA-UCL (Belgium); University of Basel (Switzerland); Universite Libre de Bruxelles (Belgium); Institut fur Mikrobiologie (Germany); CSIC/Universidad de Salamanca (Spain); IRMW-ULB (Belgium); and Katholieke Universiteit Leuven (Belgium).
The research was supported by the National Institutes of Health, the Medical Research Council of Canada, the European Commission, and the Swiss Federal Office for Education and Science.
For more information: Elizabeth A. Winzeler, Department of Biochemistry, Stanford University School of Medicine, Stanford, CA 94305-5307. Tel: 650-723-6503. Email: winzeler@cmgm.stanford.edu.
