Use of EZ:TN Transposomes for genetic analysis and direct sequencing of bacterial genomic DNA
By Les Hoffman and Jerry Jendrisak, EPICENTRE Technologies
Introduction
Methods
Results and Discussion
Conclusions
Introduction (Back to Top)
The genomes of over sixty bacterial species are sequenced or are currently undergoing sequencing. Sequencing tools have now progressed to the point at which the chromosomal DNA of small bacterial genomes can be directly sequenced without molecular cloning.1 Although direct sequencing has not been attempted for large-scale sequencing projects, the ability to directly sequence bacterial DNA has applications in gap filling and characterization of mutations. In addition, as more bacterial genome data is collected there is a need for global techniques for analyzing the functions of genes. Transposons have long been recognized as powerful tools for inserting sequencing primer binding sites and for creating gene "knockouts" (insertional mutagenesis) in microorganisms.
However, traditional transposition systems are often difficult to work with and limited in the number of microbial species in which they can be used. EZ::TN Transposomes, based on the pioneering work of Reznikoff and Goryshin,2,3 offer a new, faster, simpler and more efficient method to randomly introduce transposons into the genomes of many different microorganisms. An EZ::TN Transposome is a stable synaptic complex formed between the hyperactive Tn5 transposase and a Tn5-derived transposon. Although not required, the transposons used typically contain a selectable marker (e.g. antibiotic resistance gene) which allows rapid selection of transposed cells. Although EZ::TN Transposomes are normally formed transiently during transposition, a stable Transposome can be formed and isolated in the absence of Mg2+ (Figure 1).
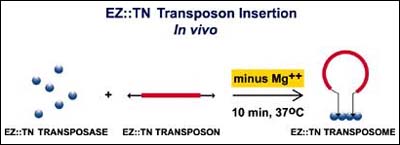
Figure 1
EZ::TN Transposomes are stable enough to be electroporated into living cells3,4 (Figure 2). Once inside the cell, the Transposome is activated, presumably by the Mg2+ within the host's cellular environment. Once activated, the EZ::TN Transposome efficiently and randomly inserts its transposon into the genomic DNA of the host cell. By screening antibiotic-selected transposition clones for a desired phenotype, gene knockouts in many open reading frames (ORFs) can be found. Thus, EZ::TN Transposomes eliminate the need for transductions or matings to mobilize transposons. In this report, we extend the use of EZ::TN Transposomes to randomly create insertion mutants in Salmonella typhimurium, Proteus vulgaris and Pseudomonas sp. Importantly, we also demonstrate that EZ::TN Transposomes facilitate direct DNA sequencing of mutated bacterial genomic DNA.
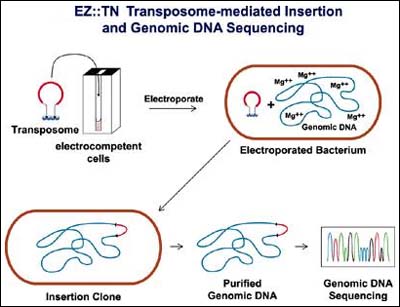
Figure 2
Transposon mutagenesis
EZ::TN
Cell electroporation
Proteus vulgaris was obtained from ATCC (number 13,315) and Pseudomonas sp. (MMSS-8 strain) was from the University of Wisconsin Bacteriology Department Stock Culture Collection. Electrocompetent cells were prepared in the same manner for all species. Cells were grown to mid-log phase at 37°C in LB broth with shaking, then chilled, harvested by centrifugation and washed with deionized water three times before suspending them in ice-cold 10% glycerol in deionized water. Cells were stored frozen at -70°C in 100-µl aliquots until used.
Prior to use, cells were thawed on ice. Fifty microliters were transferred to a 2.0-mm gap electroporation cuvette. One microliter of EZ::TN
Bacterial DNA isolation
Individual Kanr transposition clones were grown overnight at 37°C in LB Broth containing 50 µg/ml kanamycin for S. typhimurium and P. vulgaris and 300 mg/ml kanamycin for Pseudomonas sp. Genomic DNA was purified using the MasterPure Complete DNA Purification Kit (Epicentre).
Direct genomic DNA sequencing and sequence analysis
Transposon insertion sites were sequenced bidirectionally using sequencing primers specific for the ends of the inserted transposon. Two to three micrograms of bacterial genomic DNA and 5-12 pmoles of primer were used in "2X" Big Dye Terminator sequencing reactions according to the manufacturer's protocols (PE Biosystems, Foster City, CA). Bacterial genomic DNA does not require restriction endonuclease digestion or shearing to serve as a DNA sequencing template. Samples were cycled (DNA Engine, MJ Research, Waltham, MA) for 4 min at 95°C, then 60 cycles of 30 sec at 95°C and 4 min at 60°C followed by 4°C indefinitely. Sequencing reactions were purified by gel filtration with a Centri-Sep spin column (Princeton Separations, Princeton, NJ), concentrated by ethanol precipitation, washed with 70% ethanol, and resuspended in 20 ml of Template Supression Reagent (PE Biosystems, Foster City, CA). After denaturing at 95°C for 5 min, the samples were injected into an ABI 310 Genetic Analyzer (PE Biosystems, Foster City, CA) and analyzed with ABI version 3.3 sequence analysis software. Transposon insertion sites can also be sequenced using radioactive and LI-COR sequencing methods (Ronald Meis, Epicentre, unpublished data). The genomic transposition sites were located using BLAST programs maintained at the NCBI web site of the National Library of Medicine.
The primer EB-L was used for sequencing 16S rDNA.5 Cycle sequencing and sample preparation and analysis were the same as above.
Results and Discussion (Back to Top)
As demonstrated previously,3,4 EZ::TN Transposomes can be electroporated into E. coli and yeast and lead to transposon integration into chromosomal DNA. DeAngelis6 recently described a method for direct transposon-primed sequencing of bacterial genomic DNA that entails use of restriction endonuclease digestion and size selection of genomic fragments. In this report we extend the list of species successfully transposed by EZ::TN Transposome technology. We also describe simplified and improved techniques for inserting, locating and sequencing the disruption sites of introduced transposons.
EZ::TN Transposome-mediated transposon insertion
Electroporation of EZ::TN Transposomes into cells eliminates the need for matings and suicide vectors for transposon insertions into the genome of microorganisms. To demonstrate the applicability of the EZ::TN Transposome technology to a wide variety of microorganisms, we show here that Pseudomonas sp., Salmonella thyphimurium and Proteus vulgaris can be successfully and stably transposed as determined by their resistance to kanamycin (Table 1). Electroporation of EZ::TN Transposomes was found to vary in efficiency from one bacterial host to another. The variation of efficiency is most likely due to use of sub-optimal electroporation conditions.

Table 1. Average number of transposition clones generated by electroporation of EZ::TN Tnp Transposome. All values are in number of Kanr colonies per µg DNA.
One interesting finding was a readily-recognized phenotype change to one P. vulgaris transposition clone. P. vulgaris is normally a motile organism that demonstrates a "swarming" growth phenotype on agar plates. One of the P. vulgaris transposition clones was unable to swarm indicating that a gene knockout had probably been created in this clone.
Direct genomic DNA sequencing of transposon insertion sites
Genomic DNA from four P. vulgaris transposition clones, including the non-swarming clone, was purified and directly sequenced as described in Methods. Two to 2.5 µg of P. vulgaris DNA were sufficient to obtain sequence reads of approximately 400 bases (Figure 3). Using the primers from each end of the transposon, nearly a kilobase of sequence could be read from a single transposition clone.
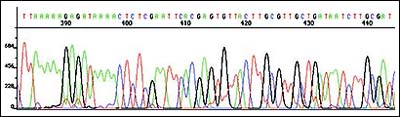
Figure 3
BLAST homology analysis of genomic sequence data generated from the four P. vulgaris transposition clones identified the specific transposon insertion site of each (Table 2). Two were identified by direct matching to P. vulgaris sequences and one by BLAST homology to an E.coli tRNA gene. The tRNAphe gene is highly conserved between E. coli and P. vulgaris, but sequence homology diverges rapidly beyond the coding region of the tRNA. Another transposon landing site was a homolog of the nifR3 gene of Rhodobacter, which is induced in response to nitrogen limitation. The third transposition clone had an interruption of a large subunit ribosomal RNA (rRNA) gene. There are probably seven or more copies of the rRNA gene in P. vulgaris
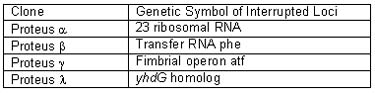

Table 2. Identification of transposon insertion sites in Proteus vulgaris transposed by EZ::TN
The transposon insertion in the non-swarming mutant clone of P. vulgaris was discovered to be within its fimbrial gene (pilus protein) homolog, atf, one of at least 45 genes involved in the Proteus swarming phenotype.7 Thus, EZ::TN Transposome disruption of the pilus protein gene would explain the loss of the swarming phenotype. Our results demonstrate rapid production of a gene knockout and correlation of cell phenotype with the genomic locus by direct genomic DNA sequencing.
Direct genomic sequencing of several randomly selected Pseudomonas sp. transposition clones led to the identification of the chromosomal locations of several of the clones. Interestingly there was no complete homology with the complete Pseudomonas aeruginosa sequence for any of six transposition clones sequenced. One explanation was that the bacterium was not P. aeruginosa as it had been designated. The sequences of the 16S ribosomal RNA genes of many bacteria are known and may be used to classify them taxonomically. We used direct genomic DNA sequencing from a 16S rDNA primer to tentatively identify the species named P. aeruginosa in the strain collection. The best match of 16S rDNA was to a polycyclic aromatic hydrocarbon-degrading Pseudomonas species originally found in creosote-contaminated soils.8 Its16S rDNA sequences are 87–95% homologous to P. aeruginosa sequences, and except for antibiotic sensitivities, other properties of the bacterium are not known.
Identifying the MMSS-8 bacterium as a non-P. aeruginosa species made the localization of transposon insertion sites more ambiguous. The strongest homologies were to the P. aeruginosa MexE gene involved in efflux systems responsible for low level multidrug resistance9 and in another clone for the P16 subunit of the MvaT transcriptional activator.10 The Mva operon is involved in isoprenoid synthesis in pseudomonads, other gram negative bacteria and mycobacteria. The synthetic pathway for isoprenoids is absent in vertebrates and man and may present a useful target for antibacterial drugs.
Conclusions (Back to Top)
EZ::TN Transposome technology is a unique, simple means of introducing any DNA segment into bacterial chromosomes. This new technology allows the in vivo insertion of transposons into several bacterial species. In fact, with improvements in the efficiencies of bacterial electroporation, there is reason to believe that in vivo Transposome-mediated mutagenesis with EZ::TN Transposomes could be applied to any bacterium for which a selectable marker exists. Not only is the EZ::TN Transposome-mediated transposon insertion process easy and fast, but the inserted transposons facilitate rapid analysis of the transposon insertion sites by such means as direct genomic DNA sequencing.
The power of EZ::TN Transposomes is further expanded by the availability of an EZ::TN pMOD
Acknowledgements
Thanks go to John Lindquist of the University of Wisconsin-Madison Department of Bacteriology for the Pseudomonas sp. culture. We are indebted to Igor Goryshin and William Reznikoff at the University of Wisconsin-Madison Department of Biochemistry for introducing us to the multitudinous possibilities lying ahead for transposome-mediated gene hopping.
Note: The EZ::TN
References
- C.R. Heiner, K.L. Hunkapiller, S.-M. Chen, J.I. Glass and E.Y. Chen, Genome Research 8: 557-561 (1998).
- I.Y. Goryshin and W.S. Reznikoff, J. Biol. Chem. 273: 7367-7374 (1998).
- W.S. Reznikoff and I.Yu. Goryshin, Epicentre Forum 6-2: 5-7 (1999).
- L.M. Hoffman, E.I Moan, J.J. Jendrisak, I.Y. Goryshin and W.S. Reznikoff , Current Genetics 35: 304 (1999).
- M.A. D. Brow, et al, J. Clin. Microbiology 34: 3129-3137 (1996).
- P.L. DeAngelis, Microbial Pathogenesis 24: 203-209 (1998).
- Allison, C. and Hughes, C. Molecular Microbiology 5: 1975-1982 (1991).
- J. G. Mueller, R. Devereux, D.L. Santavy, S.E. Lantz, S.G. Willis and P.H. Pritchard, Antonie Van Leeuwenhoek 71: 329-343 (1997).
- J.C. Pechere, M. Michea-Hamzhepour and T. Kohler, Bull. Acad. Natl. Med. 182: 599-612 (1998).
- S.R. Putra, A. Disch, J.M. Bravo and M. Rohmer, FEMS Microbiol. Lett. 164: 169-175 (1998).
